
Sungrim Seirin-Lee
PI
- Position
- Professor
- Research Field
- Mathematical Biology and Medicine, Mathematical modeling, Applied Mathematics
- ORCID
- https://orcid.org/0000-0002-8466-8037
- Personal Website
- https://ashbi.kyoto-u.ac.jp/bimed-math/
Research Overview
Mathematical elucidation of pattern formation in human developmental mechanisms and establishment of a new fusion approach linking shape and autoimmune diseases
In recent years, interdisciplinary research that applies mathematics to solve various phenomena and problems has achieved significant growth and is about to change the times. Our group is engaged in cutting-edge applied mathematics research, including the elucidation of developmental processes involved in pattern formation in the life sciences and the solution of difficult problems in intractable autoimmune diseases.
Our group is developing a method that integrates mathematical model-driven and data-driven approaches to solve the mysteries of life science that approaches the essential question, “How are we created?”. In addition, for clinical medicine, where biological experiments themselves are difficult and the methods for solving them are extremely limited, we are constructing a new fusion research method that connects clinical medicine and cell biology experiments from the concept of “shape” by using mathematical modeling with a novel idea.
Our group aims to create new concepts in the regulation of cell functions and to apply them to regenerative medicine and disease treatment by unraveling the mysteries of various biological phenomena based on “pattern and shape” as keywords. Furthermore, by discovering the universality (truth) of the life derived from mathematical models, we aim to elucidate the principles underlying human biology and to pioneer mathematical human biology.
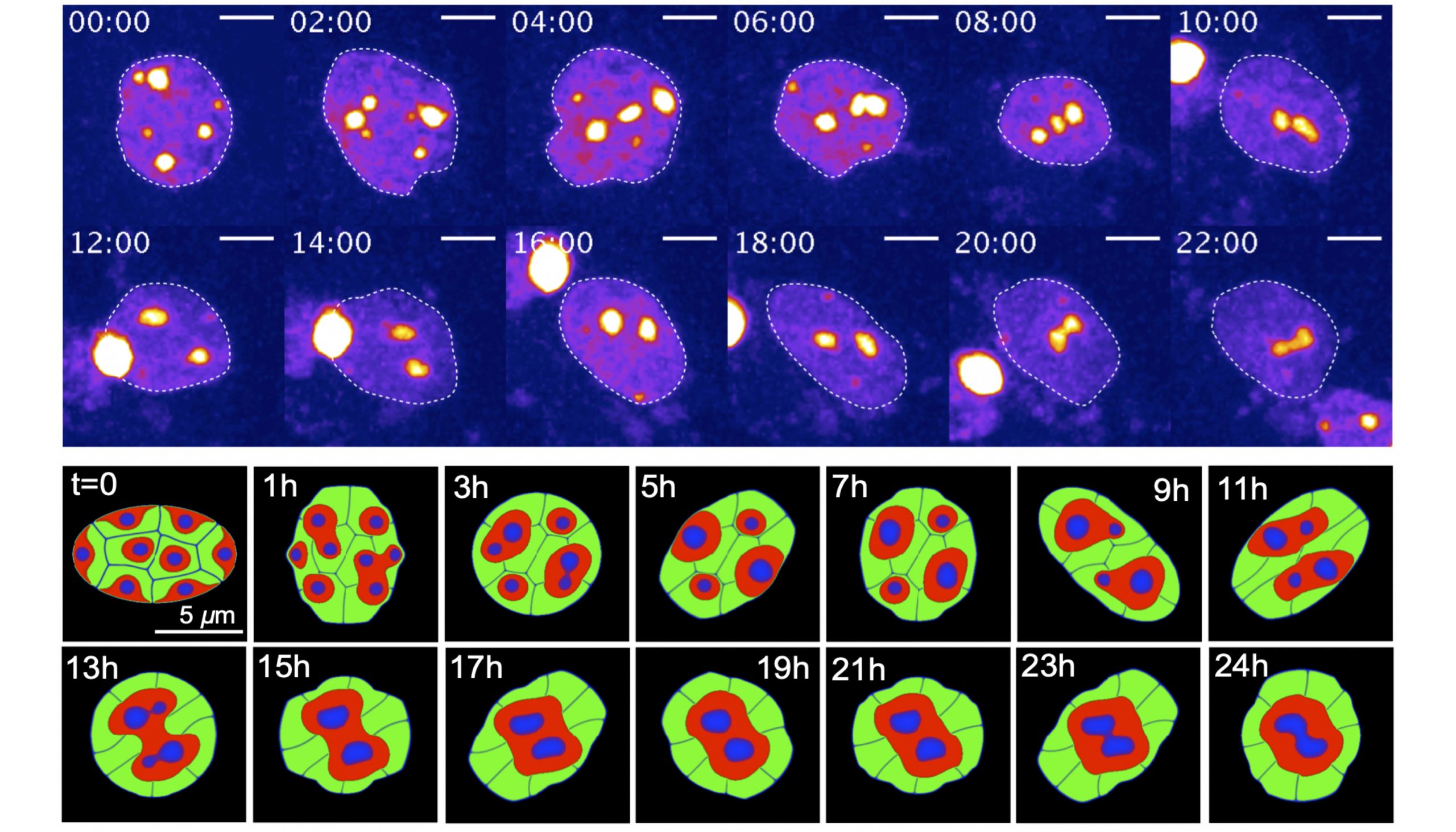
Figure 1: Dynamic nuclear deformation induces the reorganization of genomic architecture. In vitro experiment (upper panels) and the regeneration by mathematical model (lower panels).
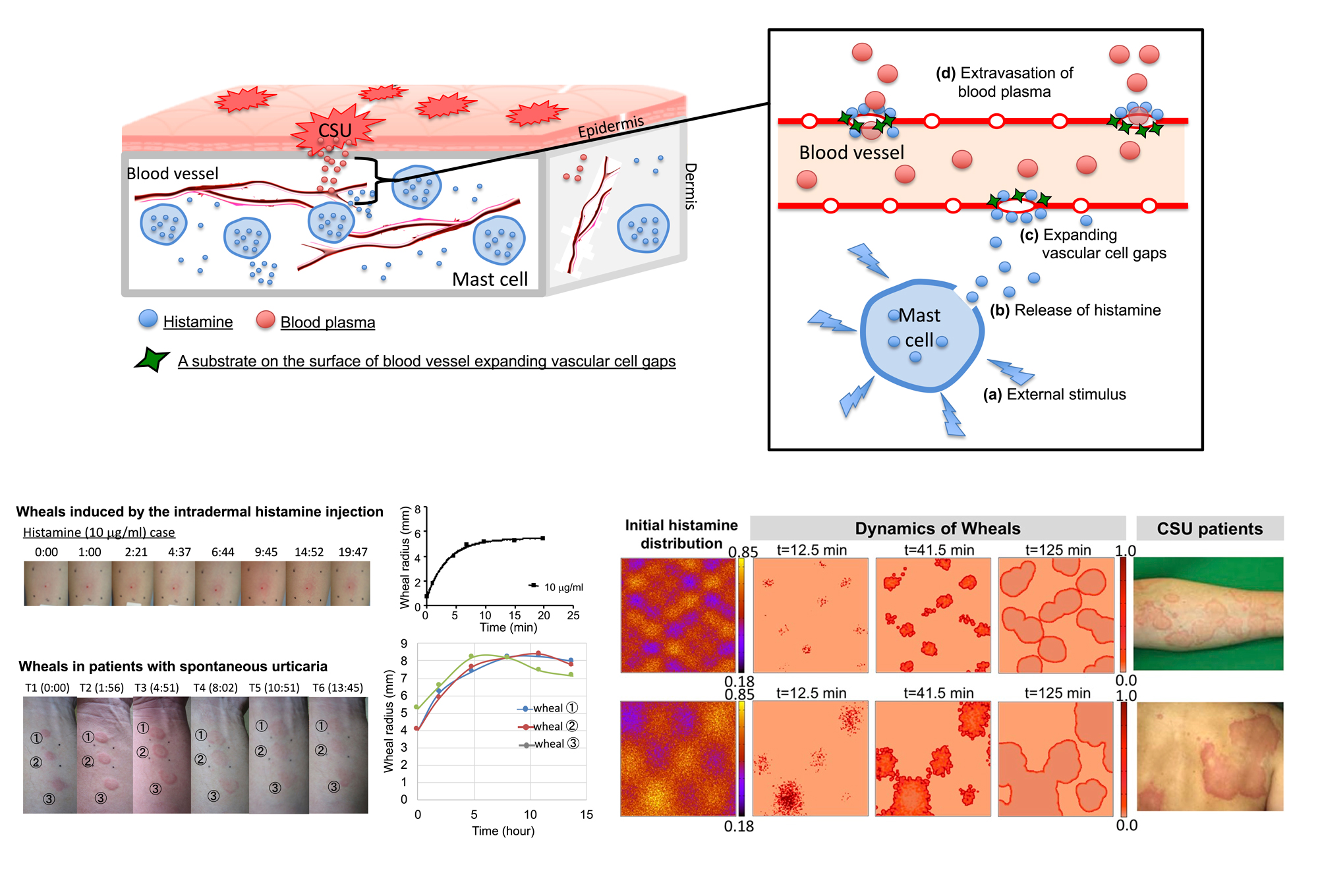
Figure 2: Chronic spontaneous urticaria (CSU). Wheal dynamics observed in patients and the regeneration of CSU pattern in skin by mathematical model.
Biography
Seirin-Lee obtained her PhD from Okayama University in 2010. During the doctoral course, she studied at Mathematical Institute, Oxford University, as a JSPS DC1 researcher. After then, she undertook postdoctoral training as a JSPS PD at the University of Tokyo and RIKEN and was appointed as an Assistant Professor in 2014, Associate Professor in 2017, Full Professor in 2020 at Hiroshima university while also working as JST PRESTO researcher from 2016 to 2019. She was appointed as a full professor at KUIAS, Kyoto University in 2021.Publications
S. Seirin-Lee*, K. Yamamoto, A. Kimura*(* Co-corresponding),The extra-embryonic space and the local contour are critical geometric constraints regulating cell arrangement (2022) Development. 149,dev200401. doi:10.1242/dev.200401 (Press Released)
S. Seirin-Lee, The role of cytoplasmic MEX-5/6 polarity in asymmetric cell division. Bulletin of Mathematical Biology. (2021) 83:29.
S. Seirin-Lee, Y. Yanase, S. Takahagi, M. Hide, Multifarious Eruptions of Urticaria Solved by A Simple Mathematical Equation. PLOS Computational Biology (2020) 16(1): e1007590. (Press Release.)
S. Seirin-Lee, M. Nomata, M. Mukunoki, Mathematical modeling and regionality-based optimal policy to reduce empty houses, Akiya, in Japan. Japan Journal of Industrial and Applied Mathematics (2020) 37:365-382
S. Seirin-Lee, Fumitaka Osakada, Junichi Takeda, Satoshi Tashiro, Ryo Kobayashi, Takashi Yamamoto, Hiroshi Ochiai, Role of dynamic nuclear deformation on genomic architecture reorganization. PLOS Computational Biology (2019) 15 (8): e1007289. (Press Release.)
M. Kuwamura, S. Seirin-Lee, S-I. Ei, Dynamics of localized unimodal patterns in reaction-diffusion systems related to cell polarization by extracellular signaling. SIAM J. on Applied Mathematics (2018) 78, No6, 3238-3257.
S. Seirin Lee, S. Tashiro, A. Awazu, R. Kobayashi, A New Application of the Phase-field Method for Understanding the Reorganization Mechanisms of Nuclear Architecture. Journal of Mathematical Biology (2017) 74, 333-354.
S. Seirin Lee, Lateral Inhibition-Induced Pattern Formation Controlled by the Size and Geometry of the Cell. Journal of Theoretical Biology (2016)404, 51-65.
Awards
MIMS Mimura Award 2020Joined
Oct. 1, 2021
