Yukihiro Yabuta
| Position | Assistant Professor |
|---|---|
| Group name | Saitou Group |
| Research Field | Bioinformatics, Developmental Biology, Molecular Biology |
| Joined | Feb. 1, 2019 |
Research Overview
Establishment of a solid basis for informatic analysis on comparative epigenetic dynamics during germ cell development in humans, Cynomolgus monkeys and mice
Germ cells undergo extensive epigenetic reprogramming/programming during their development to differentiate into gametes that contributes to next generations. Investigations into epigenetic dynamics during germ cell development are critical not only for understanding basic mechanisms regarding how our genetic/epigenetic information are transmitted, but also for diseases arising from their abnormal regulations. It is also important to understand how germ cell development are conserved across species.
In 2011, we developed a method to induce mouse ES and iPS cells into primordial germ cell-like cells (PGCLCs), which had ability to produce healthy sperm and egg and normal offsprings. Since then, we have been developing to induce PGCLCs from human iPS cells, as well as Cynomolgus monkey ES and iPSCs. By using PGCLCs, we can apply many epigenetic analysis methods, such as ChIP-seq or bisulfite-seq.
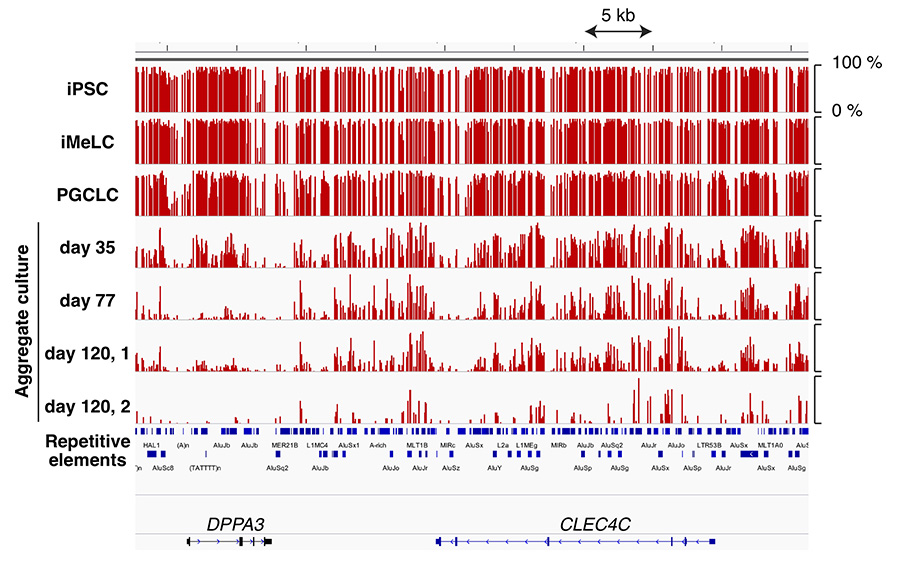
Our study showed that, PGCLCs undergo genome-wide DNA demethylation in a similar manner as the in vivo germ cells in both mouse and human. In addition, specific transposable elements were resistant to demethylation in both species. However, the demethylation time scales were different between species that, mouse PGCLCs took less than 10 days to demethylate all genomic DNAs, whereas human PGCLCs required more than 120 days. This is in a reasonable range according to the gestation period of 20 days in mouse and 280 days in human, but the regulation mechanism and difference underlying DNA demethylation are still unknown. In ASHBi, we study on how DNA methylations are regulated during germ cell development and how those are conserved between species using human, mouse and Cynomolgus monkey PGCLCs.
Biography
Yukihiro Yabuta obtained his PhD from Fukui Prefectural University (2002) and moved to U.S. and undertook postdoctoral training at Oregon Health & Science University, department of Biochemistry. He joined and worked as a postdoc at RIKEN-Kobe Institute, Center for Developmental Biology in 2003. He moved to Kyoto University, Graduate school of medicine in 2010 and worked as a postdoc. He was appointed Program-Specific Assistant Professor in 2019 in ASHBi of Kyoto University.
Publications
Nagaoka SI, Nakaki F, Miyauchi H, Nosaka Y, Ohta H, Yabuta Y, Kurimoto K, Hayashi K, Nakamura T, Yamamoto T, Saitou M. ZGLP1 is a determinant for the oogenic fate in mice. Science. 2020 Mar 6;367(6482).
Sakai Y, Nakamura T, Okamoto I, Gyobu-Motani S, Ohta H, Yabuta Y, Tsukiyama T, Iwatani C, Tsuchiya H, Ema M, Morizane A, Takahashi J, Yamamoto T, Saitou M. Induction of the germ cell fate from pluripotent stem cells in cynomolgus monkeys. Biol Reprod. 2020 Mar 13;102(3):620-638.
Yamashiro C, Sasaki K, Yabuta Y, Kojima Y, Nakamura T, Okamoto I, Yokobayashi S, Murase Y, Ishikura Y, Shirane K, Sasaki H, Yamamoto T, Saitou M. Generation of human oogonia from induced pluripotent stem cells in vitro. Science. American Association for the Advancement of Science; 2018 Oct 19;362(6412):356–60.
Mitani T, Yabuta Y, Ohta H, Nakamura T, Yamashiro C, Yamamoto T, Saitou M, Kurimoto K. Principles for the regulation of multiple developmental pathways by a versatile transcriptional factor, BLIMP1. Nucleic Acids Res. 2017 Dec 1;45(21):12152–69.
Miyauchi H, Ohta H, Nagaoka S, Nakaki F, Sasaki K, Hayashi K, Yabuta Y, Nakamura T, Yamamoto T, Saitou M. Bone morphogenetic protein and retinoic acid synergistically specify female germ-cell fate in mice. EMBO J. EMBO Press; 2017 Nov 2;36(21):3100–19.


